Parallel I/O¶
Large scale scientific applications often require the efficient storage and retrieval of large amounts of structured data. These applications, known as data intensive applications, can include big data workloads, producer consumer workloads, bulk-synchronous workloads, and machine learning applications. In addition, for large scale simulations that take days to complete, checkpointing is essential to prevent wasting valuable computing time in the event of a system or application failure. The data is typically stored in files, which are simply streams of bytes that are abstracted by the file system as single entities. The main challenge is achieving high performance and scalability while maintaining data consistency and integrity.
I/O stack¶
The I/O stack is composed of two fundamental parts: software stack and hardware stack.
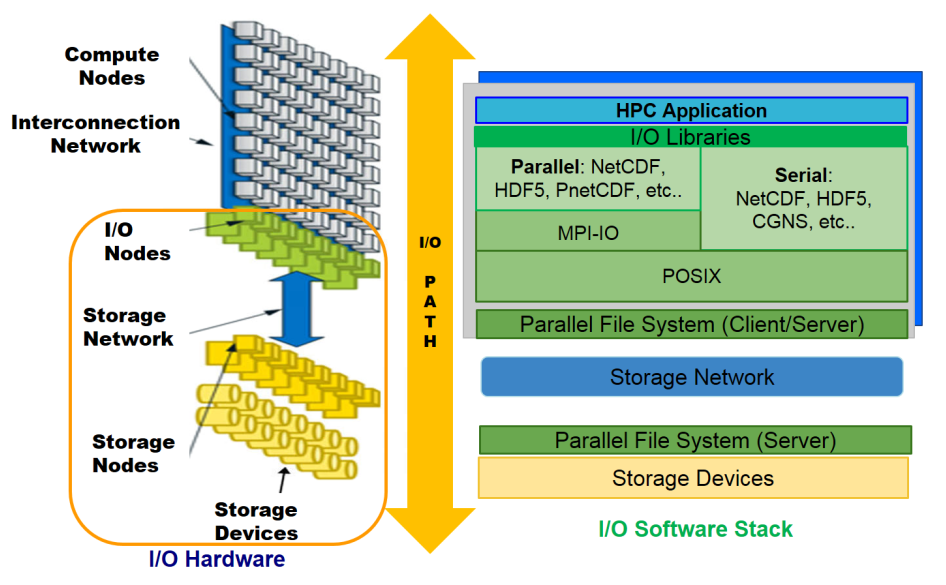
Hardware stack includes the storage nodes, the storage network, I/O nodes, and the I/O devices where data is stored physically.
Software stack includes the parallel file system which abstracts hardware complexities and provides a simple POSIX interface to the user, followed by I/O middleware that manages accesses from multiple processes. High-level libraries, built on top of the POSIX I/O and middleware, provide a more user-friendly way to store scientific data.
Scientific applications can perform I/O at all levels of the software stack
POSIX I/O¶
POSIX, or Portable Operating System Interface, is a standard that defines how the operating system accesses I/O hardware such as storage devices. A file system is considered POSIX compliant if it adheres to a set of consistency rules outlined by the IEEE POSIX standard. Parallel file systems like Lustre and GPFS are both POSIX compliant. Being POSIX compliant ensures compatibility and portability of files across different systems. However, it can impede seriously the performance of the parallel file systems by imposing consistency and atomicity requirements, adding overhead to file operations, and limiting design choices.
For these reasons, it is generally best to avoid low-level POSIX I/O in HPC applications. Instead, consider using middleware like MPI I/O and high-level libraries built on top of it.
For these reasons, depending on the I/O patterns of the HPC application it is recommended to avoid POSIX I/O.
BeeOND (BeeGFS on demand)
If your application does a lot of POSIX I/O, and integrating parallel I/O is not feasible due to the complex code base or dependencies then it may be worth considering BeeOND as an alternative.
MPI I/O¶
MPI-IO is a set of extensions to the MPI library that enable parallel high-performance I/O operations. It provides a parallel file access interface that allows multiple processes to write and read to the same file simultaneously. MPI-IO allows for efficient data transfer between processes and enables high-performance I/O operations on large datasets. It also provides additional features such as collective I/O, non-contiguous access, and file locking, which are not present in the standard POSIX I/O interface.
I/O Model¶
Data from simulations that adhere to a specific structure, such as grids or meshes, should be rearranged prior to being written to files in order to maintain this structure.
Depending on the use case, different file access models can be used:
- File per process
- Single file
- Single writer
- Independent writers
- Collective writers
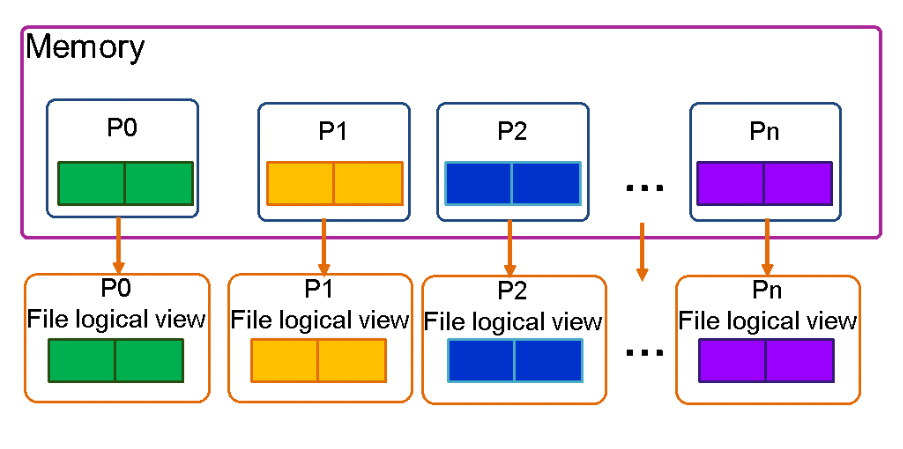
File Per Process (Multiple files per process)¶
The file per process approach is straightforward to implement as each process writes its own data to a separate file. No coordination between processes is required, making it a relatively simple implementation. Additionally, this method can achieve a high throughput rate. However, the large number of files generated can put a significant stress on the file system due to the increased number of metadata operations. Retrieving all the data for analysis can also be difficult as most analysis tools struggle to read and process thousands of files simultaneously.
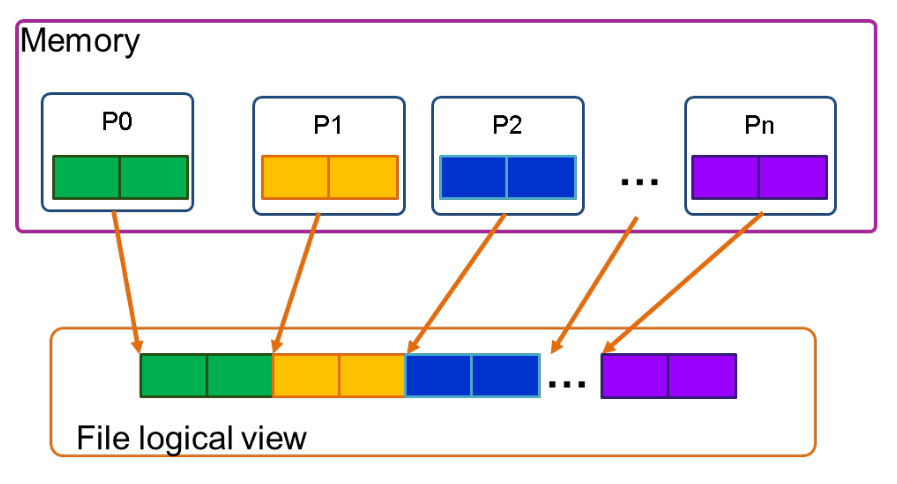
Single file¶
In contrast to the file per process method, the single file approach involves all the processes writing to a single file. This is made possible through the middleware like MPI-IO and can only be done on parallel file systems that support this type of file access.
Single writer(Serial I/O)¶
With this approach, only one process writes its own data as well as data received from other processes to a single file or multiple files. There are no benefits to using this method and it is generally not recommended, as it has two main drawbacks: it serializes the I/O step and leads to a bottleneck in communication as all processes must send data to the master process. Additionally, the master process is burdened with managing its own data and collecting data from other processes, resulting in high memory usage. Furthermore, a single node has limited I/O performance, i.e. doing I/O from multiple nodes usually provides better performance when parallel file systems are used.

Independent writers¶
With this method, all processes write to specific blocks of the file, but synchronization is necessary to prevent write conflicts. Coordination usually occurs at the parallel file system level. Although communication between processes is not required, multiple processes may compete for certain regions of the file, leading to lock contention, hence lower performance.
High-level I/O libraries such as parallel HDF5, pNetCDF, and ADIOS2 are commonly used with this approach and help to encapsulate data within the file, but the complexity of managing file regions is not visible to the programmer.
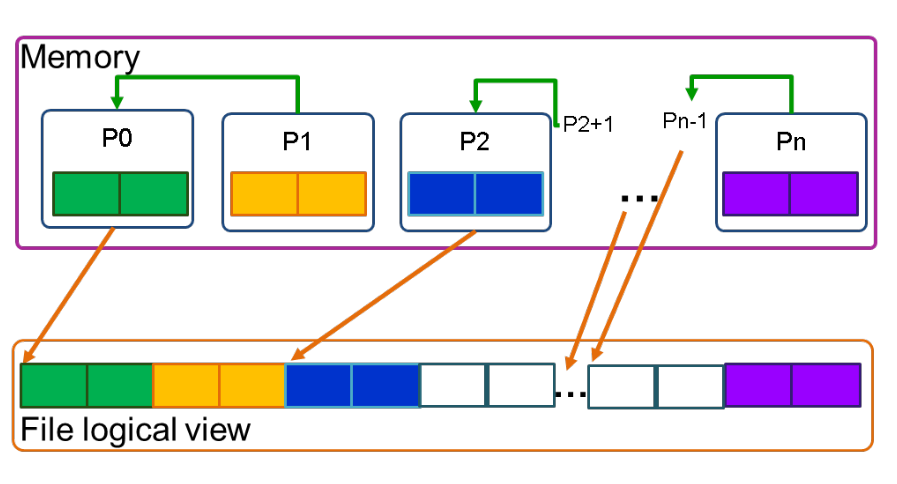
Collective writers¶
To improve performance in the single file approach, data aggregation can be done by introducing aggregators, chosen processes that collect data from others and write it to specific sections of the file. However, this method may be complex to implement.
It is worth noting that using a high-level library can make the process of data aggregation and file management much easier, as it typically handles these tasks behind the scenes and allows for a single logical view of the file while also providing control over the aggregation strategy.
Recommended resources for more details about MPI-I/O:
- High Performance Parallel I/O by Prabat & Quincey Koziol
- Using MPI 2
High level parallel I/O libraries¶
High level I/O libraries such as NetCDF, HDF5, and ADIOS2 can provide a more user-friendly way of working with structured data in scientific applications. These libraries enable representation of multi-dimensional data, labels, tags, non-contiguous data, and typed data. They also simplify visualization and analysis, and allow for data to be read and written across different machines. The following libraries are offered in HoreKA, and are accessible through the module system.
HDF5¶
Hierarchical Data Format (HDF5) is a widely used library for developing large scale scientific codes. It offers a way to organize file contents into a structure similar to a Unix-like file system, with a portable and self-describing format for datasets, attributes, data spaces, and property lists. The datasets and groups are similar to files and directories respectively.
For more details about HDF5 please refer to the documentation. Parallel HDF5 is a version of HDF5 that supports parallel access to an HDF5 file through MPI. The parallel version of HDF5 is built with MPI-I/O under the hood.
In HoreKA we offer both the serial and the parallel versions of HDF5. With the serial version only single file per process and single file single writer approaches can be applied. With the parallel version all the I/O models are applicable.
Loading the serial HDF5 module:
$ module load lib/hdf5/1.10_serial
Loading the parallel HDF5 module:
$ module load lib/hdf5/1.10
HDF5 is a feature-rich library but has a relatively steep learning curve. There are bindings available for various programming languages, including C/C++, Fortran 90, Java, Python, and R. However, parallel I/O is only supported for languages that support MPI (C/C++, Fortran, and Python). It is recommended to use high-level APIs as programming with the low-level C API can be quite challenging and bug prone.
NetCDF¶
NetCDF, short for Network Common Data Format, is a widely used library in climatology, meteorology and oceanography applications. It provides a portable and self-describing data format for storing large shared simulation data arrays. The parallel version of NetCDF, pNetCDF is parallel version of NetCDF built on top of MPI-IO.
In HoreKA we offer both the serial and the parallel version of NetCDF. With the serial version only single file per process and single file single writer approaches can be applied. With the parallel version all the I/O models are applicable.
Loading the serial NetCDF module:
$ module load lib/netcdf/4.7_serial
Loading the pNetCDF module:
$ module load lib/netcdf/4.7
NetCDF follows a hierarchical model similar to HDF5, but with fewer features and slower read/write speeds. It offers bindings for a variety of programming languages including C/C++, Fortran 77/90/95/2003, Python, Java, R and Ruby. However, similar to HDF5, parallel I/O is only supported for languages that support MPI (C/C++, Fortran, and Python).
ADIOS2¶
ADIOS2 is a powerful framework for high-performance data management, which is gaining popularity in fields such as weather forecasting, molecular dynamics simulations, and computational fluid dynamics. It offers a self-describing data format and is suitable for both efficient data storage and in-situ1 and in-transit2 visualization and analysis.
ADIOS2 engine¶
The key component of ADIOS2 is the "engine", which abstracts the data transport method. Engines can be categorized into two main groups: file-based engines and streaming engines, but most of them use the same file access API. I/O operations are performed in steps, such as time steps or iterations.
One of the key features of ADIOS2 is the ability to change the engine without having to recompile the program, by using an additional XML configuration file. This enables performance testing of different engines without requiring code modifications. Depending on the workload, different engines can be chosen, file-based engines for checkpointing and long-term data storage, SST for producer-consumer workloads, SSC for strongly coupled applications, DataMan for data movement over WAN networks, Inline for zero-copy data transfers, and NULL for testing purposes with no actual I/O performed.
Loading the adios2 module:
$ module load lib/adios2
Features of ADIOS2 that the other two libraries lack of:
- Using node local SSDs as burst buffers.
- Streaming I/O.
- Multiple data compression engines, including lossy and lossless compressions.
- GPU support, reading and writing data directly from/to host buffers.
- Implicit I/O aggregation, easily to be tuned through a single engine parameter.
- Simple High Level API for replacing the usual POSIX I/O.
- Support for data movement outside HPC sites.
- Support for multiple file formats: ADIOS2 supports multiple file formats such as its own binary file format, BP, and HDF5, allowing for more flexibility in choosing the appropriate format for a given use case. There is way more to know about the ADIOS2 library, for that you can check the documentation for more details.
Choosing the right library¶
Choosing between HDF5, pNetCDF, and ADIOS2 depends on the specific needs of your application and the requirements of your data storage and management.
- Parallel HDF5 offers a rich feature set and a hierarchical data model similar to a Unix file system, making it suitable for a wide range of use cases. However, it can have a relatively steep learning curve and may be slower for read/write operations.
- pNetCDF is similar to parallel HDF5 in terms of hierarchical data model, but it has fewer features and slower read/write speeds. It is commonly used in climatology, meteorology, and oceanography applications.
- ADIOS2 is a powerful framework for high-performance data management that is designed for extreme scale I/O. It offers a variety of engines that can be chosen based on the workload type, and the ability to change engines without recompiling the program. Additionally, it offers flexibility for in-situ and in-transit visualizations and analysis.
When choosing between the three libraries, it is important to consider the specific requirements of your application and data, as well as your experience and familiarity with each library. Additionally, it is also important to consider the community and support behind the library, as well as the compatibility with the programming languages you are using.
I/O Optimization¶
For quick tips on optimizing I/O for better throughput and metadata performance please refer to file systems performance tunning.
The main focus here will be on optimizing parallel HDF5, pNetCDF and ADIOS2.
General tuning advices:
- I/O model: Implementing the right I/O model can improve the performance drastically.
- Chunking: Improves the performance of the parallel I/O operations, as it allows for better data locality and reduces the number of metadata operations.
- Compression: Improves performance by reducing the amount of data that needs to be written to disk.
- Collective I/O: Improves the performance by reducing the amount of communication required between processes.
- Proper Buffering: Proper buffering can reduce the number of I/O operations, so that the data is written in large chunks and not in small pieces.
- Small data: Use compact storage when working will small data (<64KB).
- Rank aggregation: Tuning the aggregation settings can also improve the performance by reducing the number of I/O operations.
- Subfiling: 3 Changing the number of subfiles used by the application can result into significant performance gain. Try using more subfiles than nodes allocated. The maximum number of subfiles that can be used can't exceed the number of ranks.
Tuning parallel HDF5¶
Some tuning tips specific to parallel HDF5
- High-level API: Using high-level API-s shortens the development time and reduces the complexity of the parallel HDF5 I/O operations.
- Proper File space management: Proper file space management can reduce the number of I/O operations, and improve the performance by reducing fragmentation and making more efficient use of the file space.
- Never fill allocated chunks: Chunks are always allocated when the dataset is created. The number of allocated and initialized chunks corresponds to the number of ranks used, with more chunks more time is spent on the dataset creation phase.
- Avoid datatype conversions: Use the same datatype in memory as in the file.
Tuning pNetCDF¶
No tuning tips.
Tuning ADIOS2¶
ADIOS2 is a powerful high-performance data management framework that provides a versatile and flexible way to manage and process large amounts of data. However, to get the best performance from ADIOS2, it is important to tune and optimize its various parameters and settings. In this guide, we will cover some common techniques for tuning the performance of ADIOS2.
- Engine: It is essential to choose the appropriate engine.
- In-Situ Analysis: Analyze data while in memory, instead of writing it to disk first.
- Burst-buffer feature: ADIOS2 can take advantage of node local SSDs for I/O operations. To do that, the SSD should be specified as the data file location.
Note
For all the three libraries, the best option will depend on the specific requirements of the application and the resources of the system.
I/O Performance Analysis¶
Assessing the parallel I/O performance of a program can be challenging. Profiling and tracing tools can be used to gather information about the I/O performance without delving into the complexity of the code. A profile provides a summary of all I/O events that occur during the program's execution, while a trace provides more detailed information and includes temporal information about the I/O events. Additionally, a trace can reveal I/O performance at various levels of the I/O stack, making it easier to pinpoint performance bottlenecks.
Darshan¶
Darshan is a lightweight, scalable I/O characterization tool that transparently captures detailed information about I/O access patterns without requiring modifications to the application's source code or execution environment.
Darshan instruments the application's I/O library calls to collect information about the number of reads and writes, the size and layout of I/O buffers, and the performance characteristics of underlying storage devices.
It provides I/O profiles for C and Fortran calls including POSIX and MPI-IO, also a limited support of parallel HDF5 and pNetCDF5 is provided.
Setting up darshan¶
Darshan is made of two main components, darshan-runtime and darshan-util.
The darshan-runtime component instruments applications, the instrumentation can be done both at compile time or runtime. However, we recommend using the runtime method since only the LD_PRELOAD environment variable needs to be set, for more information please check the documentation.
On the other hand the darshan-util component is used mainly for visualizing and analyzing the logs generated by the darshan-runtime component.
In HoreKA darshan is installed with both components, to use them just load the darshan module:
$ module load compiler/gnu
$ module load mpi/openmpi
$ module load devel/darshan
GNU Compiler
It is very important to load both the gnu compiler and OpenMPI, because these were used during the darshan build.
Running an application with darshan¶
$ env LD_PRELOAD="${DARSHAN_LIB_DIR}/libdarshan.so" mpirun ...
By default the logs will be located in one of the following directories
${HOME}/Darshan-logs.${SLURM_SUBMIT_DIR}. (Path to the batch script)- PWD (Path to the executable binary)
Where YEAR,MONTHand DAY form the actual date of the application execution. From there on you can find the generated logs based on the job id,
and the execution time embedded into the logs filename.
Analyzing logs¶
There are a few options on analyzing the darshan logs:
-
Generating a quick pdf report with
darshan-job-summary.pl:$ darshan-job-summary.pl path/to/darshan-logs -
Generating a pdf report for each file accesses by the application:
$ darshan-summary-per-file.sh /path/to/darshan-logs -
Dumping all the information into a human readable ASCII file format:
$ darshan-parser --base -
For more options:
$ darshan-parser --help
Other Tools¶
Besides darshan, there are other tools that can be used for tracing and profiling I/O, like Recorder2.0 and ScoreP.
-
In situ, means both the main application and the visualization or analysis program are executed on the same machine, hence more efficient data management. ↩
-
In transit, is the opposite of in-situ, meaning that the main application and the visualization(analysis) program are running on different machines. ↩
-
Subfiling is a technique used on parallel file systems to reduce locking and contention issues when multiple compute nodes interact with the same storage node. It provides compromise between the single shared file approach that creates lock contention issues on parallel file systems and having one file per process. ↩